PerturbGen
PerturbGen is a scientific Python library for modeling, generating, and analyzing cellular perturbation effects from high-dimensional biological data. It is designed for researchers and developers working with perturbation experiments (e.g. genetic, chemical, or CRISPR-based perturbations) who want reproducible, model-driven ways to simulate and interpret perturbational responses.
Technically, PerturbGen is a trajectory-aware, generative transformer for predicting how genetic or signaling perturbations reshape transcriptional programs across cell state transitions, rather than only within a single fixed state. It formulates time, development, or lineage progression as a sequence-to-sequence (S2S) problem: given an unperturbed source cell state (and optional context from earlier observed/predicted conditions), the model predicts a target downstream state (e.g., a later time point or more differentiated fate), enabling perturbation effects introduced early or mid-trajectory to be propagated to terminal outcomes. Architecturally, PerturbGen uses an encoder–decoder Transformer: a pretrained 12-layer encoder embeds the source gene-token sequence, while a Transformer decoder (dataset-tuned depth) combines self-attention over the target sequence with cross-attention to the encoded source and contextual condition embeddings. Cells are tokenized as ranked gene sequences using a Geneformer-style tokenizer (gene ranks from normalized expression), and decoding is non-autoregressive, following a MaskGIT-style iterative refinement scheme that fills masked tokens in parallel with bidirectional context, which better matches the non-sequential nature of gene regulation than left-to-right generation. Training combines (i) a masked token prediction objective on the target condition conditioned on source + context, and (ii) an auxiliary count reconstruction head (a lightweight MLP with a probabilistic count model) to map learned embeddings back to gene expression space for interpretable transcript-level perturbation readouts. At inference, PerturbGen runs paired forward passes (control vs perturbed) where perturbations are simulated by editing the source gene-token sequence (e.g., KO by removing/zeroing a token; activation by rank promotion), producing predicted downstream expression changes and embedding shifts that can be scaled to genome-wide in silico perturbation atlases across trajectories.
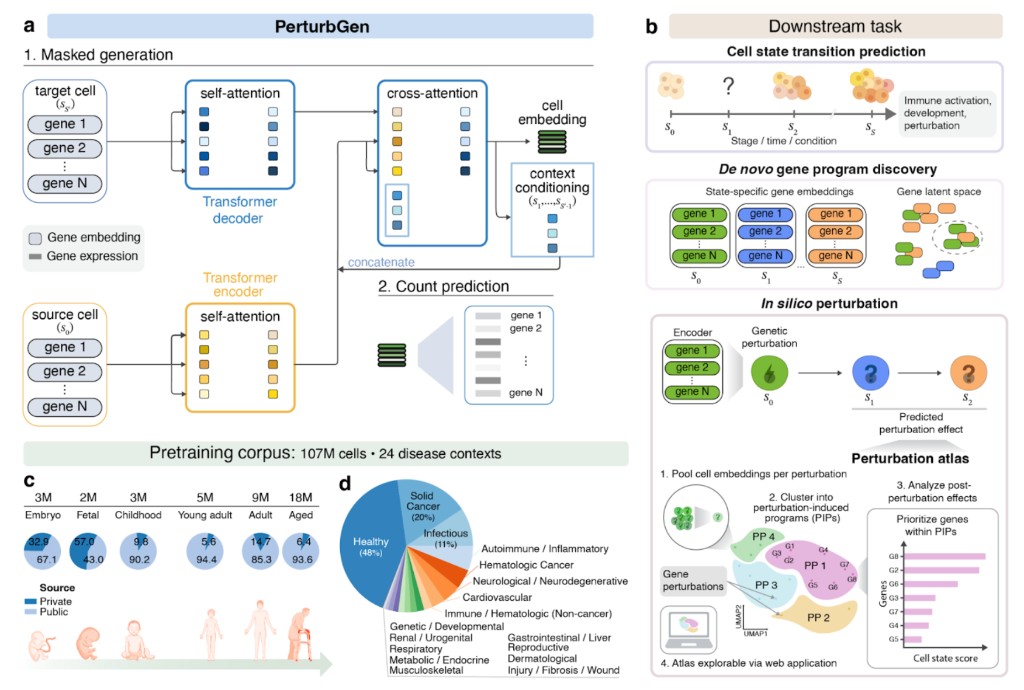
Conceptual overview
At a high level, a typical PerturbGen workflow looks like:
Prepare perturbation-aware biological data (e.g. single-cell expression)
Configure or load a perturbation model
Apply perturbations in silico to generate predicted responses
Analyze, compare, or visualize the resulting perturbation effects
PerturbGen does not aim to be a general-purpose machine learning framework. Instead, it provides abstractions that encode biological perturbations as first-class objects, making downstream analyses easier to reason about and reproduce.
See Also
- Installation
- Data
- Tutorial
- API