6. Perturbation (in silico)
This notebook runs in silico perturbation using a count-decoder checkpoint from notebook 3 (i.e. train Perturbgen) and a YAML config. Update paths in the config before running.
In the LPS time-course dataset, we first used PerturbGen’s time-resolved gene embeddings to identify transcriptional programs that change over the trajectory. Clustering embeddings across time highlighted a cytokine/chemokine program that was strongly induced early after LPS, with IL1B emerging as a central, highly responsive gene in myeloid lineages (e.g., CD14⁺/CD16⁺ monocytes and dendritic cells). See notebook 5 for GP dicovery.
from IPython.display import Image, display
display(Image(filename="../_static/fig2.png"))
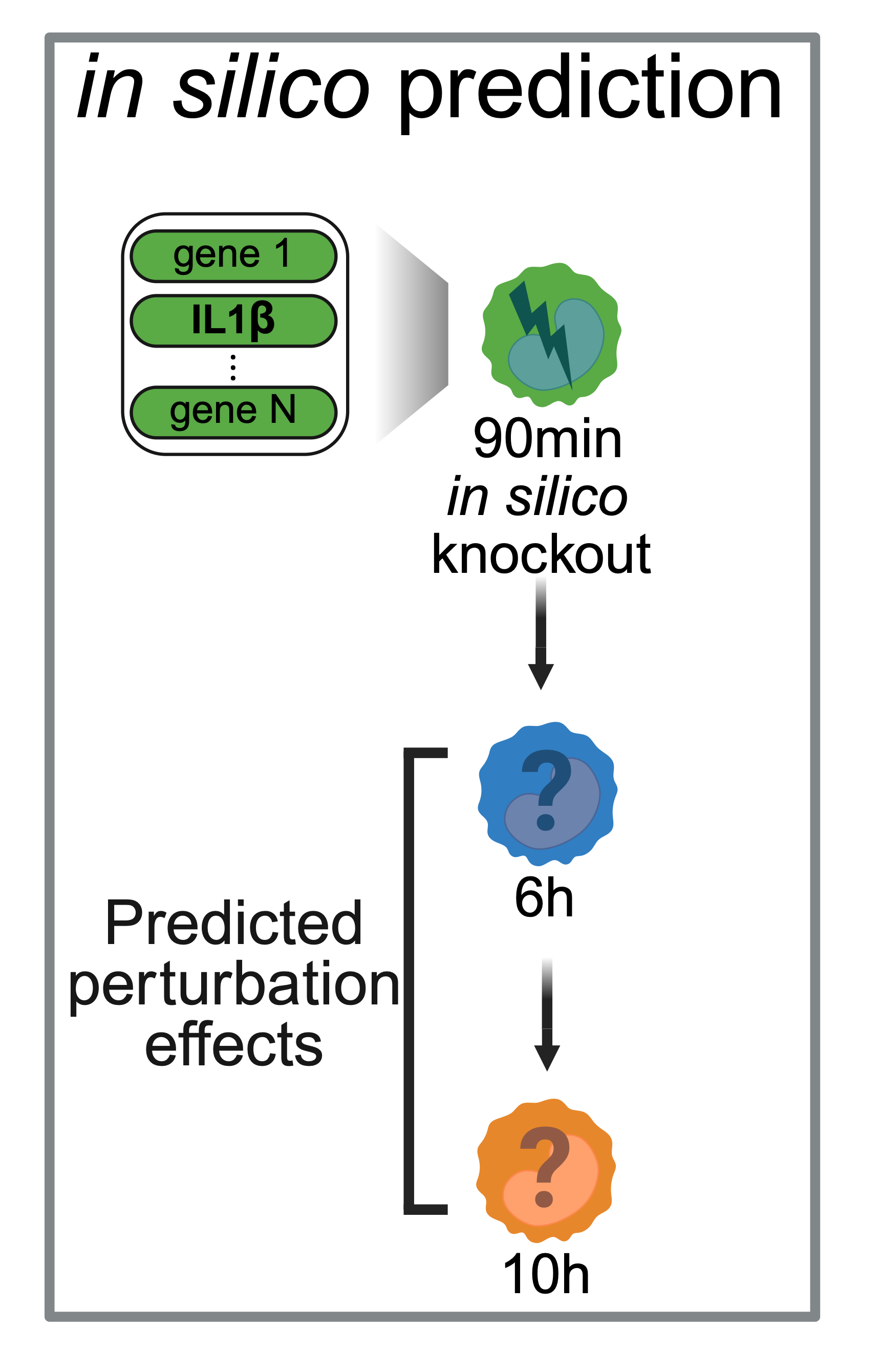
Based on this, we chose IL1B as a mechanistic candidate to test whether an early inflammatory driver can propagate effects downstream along the response trajectory. Following the schematic in Fig. above, we then performed an in silico IL1B knockout at the 90-minute time point (i.e., editing the 90-min source state), and used PerturbGen to predict the downstream transcriptional consequences at 6 hours and 10 hours. Concretely, we run two forward passes—control vs. IL1B-perturbed—starting from matched 90-min cells (with the same conditioning context), and quantify the perturbation effect at 6 h and 10 h by comparing predicted expression profiles (e.g., DEGs / pathway shifts) between the two predicted futures.
6.1. Config
Edit the config file:
docs/examples/configs/perturbation.yaml
PerturbGen perturbations are defined through a lightweight YAML configuration file, with a ready-to-use template provided in the /config folder. This design allows users to run reproducible in silico perturbation experiments without modifying model code. The key parameter controlling perturbations is:
genes_to_perturb:
- "ENSG00000125538"
This field specifies which gene(s) are perturbed in the source or specified time state. Each entry can be provided as either: • an Ensembl gene ID (recommended for unambiguous mapping), or • a gene symbol (e.g. “IL1B”).
In the current example, “ENSG00000125538” corresponds to IL1B, which we perturb at the 90-minute time point to assess how early disruption of IL1 signaling propagates to later stages of the immune response (6 h and 10 h).
Importantly, genes_to_perturb accepts a list, enabling simulation of: • single-gene perturbations (e.g. IL1B alone), or • multi-gene perturbations (e.g. combinatorial knockouts or pathway-level interventions).
In addition to specifying which genes to perturb, the configuration template allows users to control where along the sequence the perturbation is applied via the perturbation_sequence field:
perturbation_sequence:
- "src"
This parameter defines which sequence is edited before prediction, and can take one of two values: • “src” (source perturbation) The perturbation is applied to the source state (e.g. the normal or 90-minute LPS condition depending on how you defined your source in the tokenization step). The model then predicts how this early intervention propagates to downstream target states (e.g. 6 h and 10 h). This is the default and biologically most relevant setting for studying causal, trajectory-level effects, such as early signaling disruptions that reshape later cell states. • “tgt” (target perturbation) The perturbation is applied directly to the target state being predicted. This setting is useful for within-state perturbation analysis or for comparisons with models that do not explicitly model state transitions.
PerturbGen separates where a perturbation is applied from which time points are predicted, allowing flexible experimental designs along a trajectory.
Predicting downstream time points The pred_tps field specifies which time points the model should generate:
pred_tps:
- 1
- 2
These indices refer to the ordered time points defined for the dataset. In the LPS experiment, this corresponds to: 0 → 90 minutes 1 → 6 hours 2 → 10 hours. In this example 90 min has been considered as source. So, it instructs PerturbGen to predict transcriptional states at 6 h and 10 h, using the earlier 90-minute state as conditioning input. Both predictions are produced for control and perturbed runs, enabling direct comparison of downstream effects.
When perturbation_sequence is set to “tgt”, an additional parameter becomes relevant:
pert_tps:
- 1
This field specifies which target time point(s) are perturbed, rather than the source. For example, with:
perturbation_sequence:
- "tgt"
pert_tps:
- 1
the perturbation would be applied directly at time point 1 (6 h) before predicting the corresponding output. (if source has been selected as 90m and not normal).
Importantly, pert_tps is ignored when perturbation_sequence = “src”, since source perturbations are applied upstream (e.g. at 90 minutes) and automatically propagated to all predicted downstream time points listed in pred_tps.
PerturbGen supports multiple perturbation modes, which control how the specified gene tokens are edited in the sequence. This is configured via:
perturbation_mode:
- "mask"
The following modes are supported:
["mask", "pad", "delete", "overexpress"]
Each mode corresponds to a different biological interpretation:
mask: The gene token is replaced with a[MASK]token. This simulates a loss-of-function (knockout/knockdown) by removing gene-specific information while allowing the model to infer downstream consequences from context. This is the default and recommended mode for most in silico KO experiments.pad: The gene token is replaced with a[PAD]token, effectively removing it from the active sequence. This also represents a knockout-like perturbation, but with a stronger assumption that the gene is entirely absent from the expressed gene set.delete: The gene token is completely removed from the sequence. This enforces the strongest form of gene deletion, reducing sequence length and eliminating any residual representation of the gene. This mode is useful for stress-testing robustness of perturbation effects.overexpress: The gene token is moved to the top rank of the gene sequence, simulating gene overexpression or activation (e.g., CRISPRa). Because the tokenizer encodes relative expression via rank, promoting a gene to the highest rank increases its inferred regulatory influence on downstream states.
In practice:
mask,pad, anddeletesimulate gene knockout or inhibition (CRISPR KO / CRISPRi-like scenarios).overexpresssimulates gene activation or overexpression (CRISPRa-like scenarios).
6.2. Perturbation (GPU required)
Run in silico perturbation (prediction) using val.py and a YAML config.
This uses the repository script at perturbgen/Perturb/val.py.
VAL_SCRIPT = "perturbgen/Perturb/val.py"
CONFIG_PATH = "docs/examples/configs/perturbation.yaml"
cmd = [
"python", VAL_SCRIPT,
"--config", CONFIG_PATH,
]
print(" ".join(cmd))
python perturbgen/Perturb/val.py --config docs/examples/configs/perturbation.yaml
import subprocess
subprocess.run(cmd, check=True)